In honor of Thomas Paine and other Founders & Immigrants. In memory of my daddy Horst Bingel and my mom Irma Bingel
Tuesday, April 28, 2026
Research improves molecular probe of protein binding sites for drug discovery
Monday, April 20, 2026
Discovery could lead to new therapies for blood disorders
Thursday, March 26, 2026
How huntingtin proteins travel in the brain
Amazing stuff!
"Mutant huntingtin protein, which causes the neurodegenerative disorder Huntington’s disease, travels through the brain using tiny ‘tunneling nanotubes’, research has revealed. Researchers found that a protein called Rhes binds with a protein for cellular acidity regulation, SLC4A7, to build tube-like structures creating a highway to shuttle huntingtin from neuron to neuron. Interrupting this pathway minimised the spread of huntingtin in the brain and offers a potential druggable target."
From the abstract:
"Tunneling nanotubes (TNTs) are membranous structures that mediate intercellular transfer of proteins, including the pathogenic mutant Huntingtin (mHTT) protein in Huntington disease (HD).
We previously identified the ras homolog enriched in the striatum (Rhes) as a key regulator of TNT formation and mHTT transmission; however, the molecular components underlying this process remained unknown.
Here, using unbiased liquid chromatography–tandem mass spectrometry analysis of membrane-associated Rhes complexes, we identify Slc4a7 (solute carrier family 4 member 7), an intracellular pH sensor, as a top membrane-binding partner of Rhes.
Functional studies revealed that small interfering RNA–mediated depletion or pharmacological inhibition of Slc4a7 substantially reduced Rhes-induced TNT formation and suppressed mHTT intercellular transfer.
Mechanistically, Rhes directly interacts with Slc4a7 through both its amino- and carboxyl-terminal domains and modulates intracellular pH to facilitate TNT formation. This interaction does not depend on the transporter activity of Slc4a7. However, inhibition of Rhes farnesylation—a lipid modification that anchors Rhes to the membrane—disrupts its binding to Slc4a7 and abolishes TNT formation.
Slc4a7 knock-out mice showed markedly reduced cell-to-cell transmission of mHTT in the striatum in vivo.
Together, these findings uncover a previously unrecognized Rhes-Slc4a7 signaling axis critical for TNT-mediated mHTT transmission and highlight Slc4a7 as a potential therapeutic target to limit disease spread in HD."
Targeting Tunneling Nanotubes Reduces Spread of Mutant Huntington’s Protein
Membrane-associated Rhes-Slc4a7 complex orchestrates tunneling nanotube formation and mutant Huntingtin spread (open access)
Fig. 1. Membrane-anchored Rhes drives the formation of TNTs between cells.
Wednesday, February 04, 2026
Aging slows breakdown of synaptic proteins, raising disease risk
- A new study reveals mechanisms linking synapse loss to cognitive decline and dementia in aging brains, highlighting critical changes during this process.
- Researchers found that aging slows the breakdown of synaptic proteins, leading to accumulation that may contribute to diseases like Alzheimer’s.
- Stanford’s innovative tagging method could enable tracking of neuronal proteins, facilitating the identification of new biomarkers for assessing brain health.
Tuesday, January 06, 2026
Machine learning reveals hidden dimensions of functional similarity in proteins
Saturday, December 06, 2025
Protein-prediction tools based on AI (AlphaFold family) revolutionize science
Thursday, December 04, 2025
A tiny protein complex controls fat cell size and lipid storage
Saturday, November 29, 2025
New biosensor technology maps enzyme mystery inside cells
Thursday, November 06, 2025
Study finds targets for a new tuberculosis vaccine
Saturday, October 25, 2025
Anthrogen Introduces Odyssey: A 102B Parameter Protein Language Model
Sunday, September 28, 2025
On SimpleFold: Folding Proteins is Simpler than You Think
Tuesday, September 09, 2025
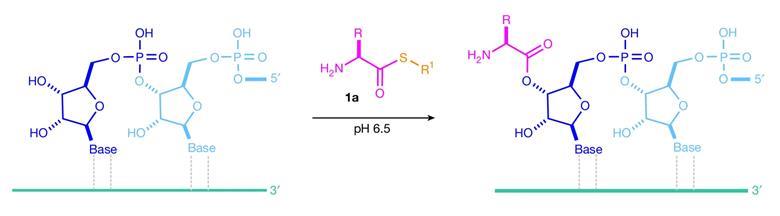
Thioesters could explain how proteins first formed on early Earth
Thursday, August 28, 2025
Scientists build an “evolution engine” to rapidly reprogram proteins
Wednesday, July 16, 2025
More than 200 misfolded proteins, much more than previously known, may contribute to Alzheimer's and dementia
- More than 200 types of misshapen proteins in the brain could be associated with age-related cognitive decline in rats.
- Researchers believe these proteins slip past a surveillance system in the cell that specifically targets and destroys malfunctioning proteins.
- A better understanding of these protein deformities in the brain could lead to better treatments and preventative measures.